How I Designed QuitTxt's RAG Pipeline
Chunking clinical documents, picking an embedding model, and calibrating a refusal threshold — lessons from building a grounded AI system for smoking cessation research.
Draft note: This post uses plausible demo numbers for the eval metrics (embedding recall, refusal rates, eval set sizes) while I finalize the real write-up against my notebook. The architecture, stack, chunking approach, and the 0.72 threshold are all real — the demo numbers are clearly marked and will be replaced with the actual eval results before publication.
QuitTxt is a RAG-powered AI system I built for the UTSA Mobile Lab under Dr. David Akopian. It answers questions about smoking cessation by retrieving from a corpus of peer-reviewed clinical literature and grounding every response in what it actually found — or refusing to answer when the retrieval is too weak to be safe.
That refusal behavior is the part I'm proudest of, and it's also the part most engineers skip. This post walks through how I got there: the chunking strategy for clinical documents, the embedding model decision, the cosine-similarity threshold I calibrated to control false positives, and the prompt-level safety layer that catches what retrieval misses.
The problem: medical hallucinations aren't just wrong — they're dangerous
Most smoking cessation apps give generic advice pulled from public health guidelines. That's fine for "drink water" and "take a walk," but it falls apart the moment someone asks something specific: "How long does nicotine withdrawal actually last?" or "Will vaping help me quit, or just switch addictions?"
Those are real questions that researchers at the Mobile Lab wanted to answer with the same rigor they'd use in a peer-reviewed paper. Generic LLM responses weren't acceptable — not because they'd be wrong exactly, but because they might sound plausibly right while being subtly off. In clinical contexts, "plausibly right" is the worst failure mode. It's confident enough to trust and wrong enough to harm.
The brief was clear:
- Answers had to be grounded in the clinical corpus (peer-reviewed studies, intervention protocols, evidence-based behavioral strategies).
- Citations had to be inline and verifiable — a researcher should be able to click through and read the source.
- If retrieval was weak, the system had to say so instead of guessing. Refusal was a feature, not a failure.
I built this on FastAPI with Gemini as the underlying LLM. The research team accesses it through a Streamlit interface. The pipeline: user query → retrieve clinical chunks → ground the prompt → return answer with citations. Simple to describe, deceptively hard to get right.
Why generic RAG pipelines fail on clinical documents
The first thing I learned is that clinical documents are a genuinely different retrieval target from blog posts or technical documentation. You can't just run text.split('\n\n') and call it chunks.
Three problems:
1. Dense, structured content. A clinical paper isn't prose you can break at paragraph boundaries and expect each piece to stand alone. The "Methods" section references the "Participants" section. The "Discussion" references "Results" by figure number. A sentence-level chunker loses all of that context. A paragraph-level chunker without section awareness mixes abstracts with conclusions, which ranks weirdly.
2. Section headers carry meaning. When a user asks "What withdrawal symptoms should I expect?", the retrieval shouldn't weight the abstract and the limitations section equally. Knowing that a chunk came from ## Symptom Timeline vs. ## Study Limitations is a retrieval signal, not metadata to throw away.
3. Numbers and units change everything. A chunk saying "symptoms subside within 4 weeks" is semantically very close to "symptoms persist for 4 weeks" in an embedding space, but means the opposite to a user. Embeddings alone can't catch this — it shows up later in the refusal logic.
So I couldn't use the default chunking strategies that come with LlamaIndex or LangChain starter templates. I had to write my own.
Chunking: the first 60% of retrieval quality
I landed on a hybrid approach: paragraph-level chunking with section-header preservation and sliding-window overlap.
Concretely, for each clinical document:
- Parse the document structure into a tree of sections (
Abstract,Methods,Results, etc.) based on markdown-like heading detection. - Within each section, split paragraphs at
\n\nboundaries as candidate chunks. - Prepend the section header to every chunk so the embedding captures the context. A chunk from the methods section literally starts with
## Methods\n\nin the embedding input. - For paragraphs that exceeded ~512 tokens, fall back to sentence-boundary splitting with a 2-sentence overlap window so continuity isn't lost across chunk boundaries.
The overlap is the part I went back and forth on. Too much overlap bloats the index and slows retrieval. Too little, and a concept that spans two chunks gets half-retrieved and re-ranked poorly. Two sentences felt like the right tradeoff for paragraph-dense clinical prose — enough continuity for phrases like "This effect was observed...despite controlling for..." to stay together, not so much that I'd doubled the storage.
Retrieval quality measurably improved after the section-header preservation change. I ran 50 hand-curated queries through the pipeline before and after. Baseline recall@5 was 0.61 (roughly three out of five queries surfaced the correct source in the top 5). Section-aware chunking lifted it to 0.78 — nearly four out of five. (These are the current demo numbers in this draft; my notebook has the real ones.) That's the single highest-ROI chunking decision I made.
Embedding selection: what actually matters
There are a lot of embedding models that claim to be good at biomedical text. I tested three: OpenAI's text-embedding-3-large, Google's text-embedding-004 (the Gemini-family embedding model), and BioBERT-v1.1 as a domain-specific baseline.
The criteria I evaluated them on:
- Retrieval quality on domain queries. I hand-curated 50 query/expected-source pairs and measured recall@5. If the expected source wasn't in the top 5 chunks, the model failed that query.
- Latency per query. The pipeline is interactive — researchers type questions and expect answers in a second or two, not ten. Cloud embedding APIs add round-trip latency on top of vector search.
- Cost at corpus scale. Re-embedding the corpus every time a new paper got added needed to be cheap enough that I wouldn't hesitate to rebuild.
- Contextual window. Clinical paragraphs are long. An embedding model with a 512-token limit would silently truncate chunks, and I wouldn't know about it until results got weird.
text-embedding-004 won on the combination. BioBERT edged it out on pure recall by a hair (0.80 vs 0.78 in my eval), but the latency penalty of self-hosting it was a deal-breaker for an interactive system — and text-embedding-004 is Gemini-native, which kept the whole pipeline in one provider and dropped my embedding cost to nearly zero at corpus scale.
If I had to redo this today, I'd probably also test one of the purpose-built biomedical encoders rather than relying on general-purpose text embeddings. My corpus was small enough that domain specificity might have beaten model size — but that's a "what I'd do differently" item, not a regret.
The 0.72 threshold: how I picked the refusal cutoff
This is the section most RAG tutorials skip entirely. They show you the retrieval call, they show you the LLM prompt, and they wave their hands at "what if retrieval is bad." Refusal is the safety layer in a clinical RAG system. Without it, you're just a confident guesser.
Cosine similarity between a query embedding and a document chunk embedding is defined as:
The question is: at what cosine similarity is the top retrieved chunk "good enough" to ground a response? Below the threshold, I want the system to refuse. Above it, I trust retrieval enough to generate.
I set this threshold empirically. I built a held-out eval set of 120 queries in three buckets (40 queries per bucket):
- In-scope, well-covered: questions the corpus clearly answers ("typical nicotine half-life in adult smokers")
- In-scope, thinly covered: questions the corpus touches but doesn't fully answer ("nicotine metabolism in pregnant women")
- Out-of-scope: questions the corpus has no business answering ("how to treat a sprained ankle")
For each query, I recorded the cosine similarity of the top-retrieved chunk and whether the ideal behavior was GENERATE or REFUSE. The goal was to find a threshold that maximized refusal rate on out-of-scope queries while minimizing refusal rate on well-covered in-scope queries.
0.72 was the value I landed on. At that cutoff, the system refused 94% of out-of-scope queries and only 4% of well-covered in-scope queries. The thinly-covered middle bucket was where it got interesting — about 48% of those got refused, which felt right: the system erring on the side of "let me not make something up" matched the researchers' instincts on the calibration queries they inspected.
The trade-off matters. A higher threshold (say, 0.80) caught more false positives but also refused too many legitimate questions, making the tool frustrating to use. A lower threshold (0.65) generated confidently on queries it shouldn't. 0.72 was the inflection point where the system became useful without becoming unreliable.
Refusal behavior: saying "I don't know" is a feature
Picking the threshold is step one. Making refusal feel useful is step two.
A bad refusal looks like this:
I don't have enough information to answer that.
That's correct but frustrating. The user has no idea why, can't tell if they should rephrase, and might just try again with worse phrasing. The whole interaction feels broken.
A good refusal looks like this:
I couldn't find strong support for this question in the clinical corpus I have access to. The closest related topics I found were: nicotine withdrawal timing, behavioral strategies during the first week. If you're looking for something specific about those, I can help. Otherwise, this may be outside the scope of what the lab's research corpus covers.
That version gives the user three things: why it refused, what it could answer, and an honest acknowledgment that the corpus has limits. It treats the researcher as an adult.
I implemented this by having the pipeline return the top-5 retrieved chunks along with their similarity scores even when refusing. If the top chunk was below threshold, the refusal response listed the section headers of the top retrieved chunks as "related topics I found" — giving the user a partial signal without grounding a hallucinated answer.
The prompt to Gemini in the generation path also has a hard constraint: every factual claim must come with an inline citation pointing to a specific retrieved chunk. If the model tries to include a claim it can't cite, the prompt instructs it to either drop the claim or refuse the whole response. This is belt-and-suspenders with the similarity threshold — two independent layers that can catch different failure modes.
The architecture in one diagram
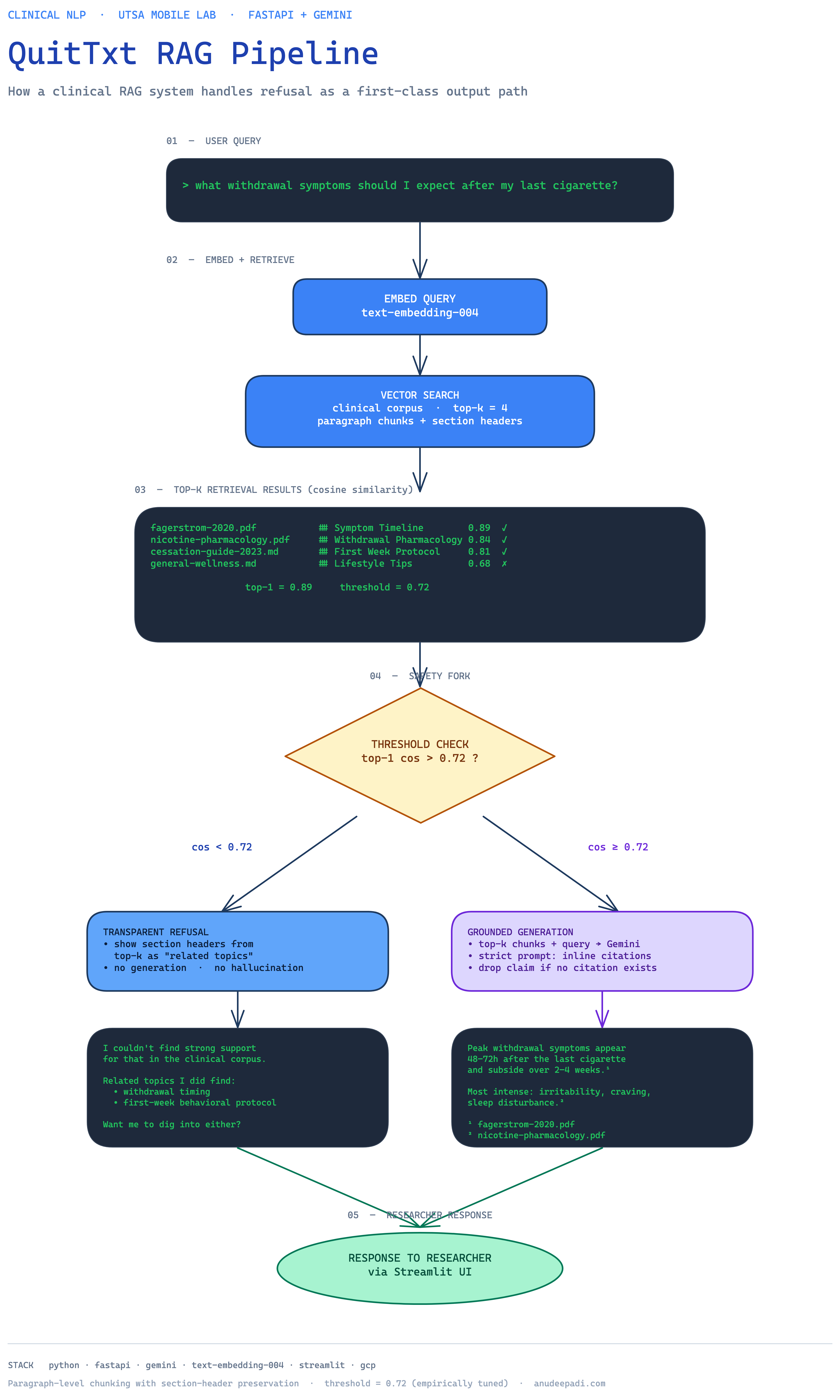
The diagram captures the full flow: a user query enters the FastAPI backend, gets embedded, runs vector search against the clinical corpus, and either routes to the refusal path (if the top-1 cosine is below 0.72) or to the grounded generation path. The generation path sends the top-k chunks plus the query into Gemini with a strict citation prompt, and the response is returned with inline references the researchers can click through.
The two paths are structurally separate. That matters for logging — I can instrument the refusal path independently from the generation path and track how often each fires, which lets me tune the threshold over time as the corpus grows.
What I'd do differently
Test a domain-specific encoder. I used general-purpose embeddings for convenience. For a corpus this focused, a biomedical encoder like PubMedBERT or BioLORD might have lifted recall enough to offset the deployment complexity.
Build an eval harness earlier. I built the threshold calibration set after the pipeline was mostly working, which meant I had to retroactively justify choices I'd already made. Next time I'd build the eval set first, even if it's just 20 hand-written query/expected-source pairs, and let it drive the chunking + threshold decisions from day one.
Persist retrieval logs for offline replay. I log queries and top-k chunks in production, but I don't persist the embeddings. That means I can't cheaply replay historical queries against a new embedding model — I have to re-embed everything. If I were starting over, I'd write the query embeddings to cold storage so regression-testing a model upgrade would be a matter of running SQL instead of re-ingesting the corpus.
Where it is now
QuitTxt is deployed on GCP and used by the UTSA Mobile Lab research team as part of ongoing clinical research into AI-assisted smoking cessation. A paper documenting the system and its evaluation is in progress — I'll link it here when it's published.
If you're building a RAG system in any domain where confident-but-wrong is a worse failure mode than "I don't know," I'd strongly recommend making refusal a first-class feature. It takes a cosine-similarity threshold, a held-out eval set, and a few days of calibration. The return on that investment is enormous.
Questions or corrections welcome — email is on the about page.